3.Octopus-toolkit output directory¶
Octopus-toolkit creates five directories when you run the program.
- Octopus-toolkit main directory.
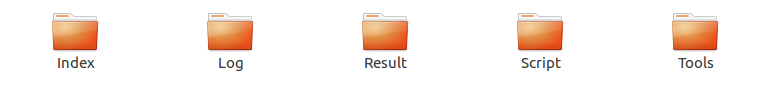
| Main Folder | Sub Folder | Description |
|---|---|---|
Index |
Reference, Hisat2, STAR | Reference genome sequence and annotation files for analysis and alignment tools. |
Log |
Command, Run | Log file containing the commands used for analysis. |
Result |
GSE_Folder, P_Folder | The output files. |
Script |
Scripts used for analysis. | |
Tools |
Analysis tools | Store the 3rd party tools used by Octopus-toolkit. |
3-1.Index-Reference¶
Reference folder
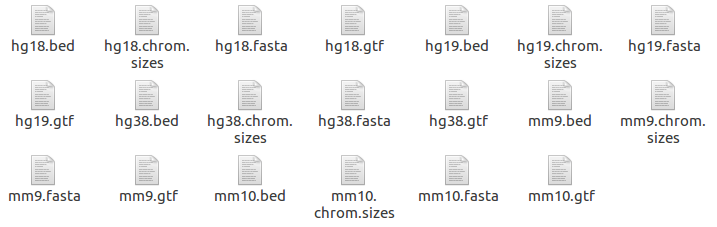
The reference folder contains several reference files required for analysis.
Before starting each process, Octopus-toolkit checks the folder whether the reference files are prepared. If not, it automatically prepares the files.
3-2.Index-Hisat2¶
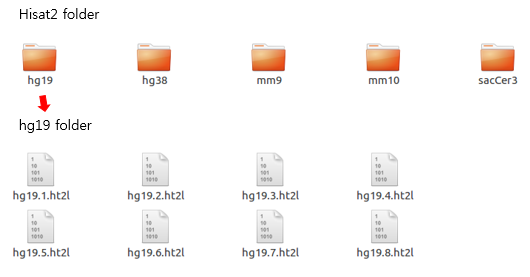
The reference genome sequence file should be indexed at least once before proceeding to the alignment step. The folder contains indexed genome sequence files used by the Hisat2 tool.
Octopus-toolkit inspects the index file of the genome before running the alignment process and runs the indexing step if it does not exist.
3-3.Log-Command¶
Command folder
The Command directory contains log files containing the commands used during the analysis.
The file name adopts the date it is created. (2016_Dec_06.cmd.txt)
3-4.Log-Run¶
Run folder
The Run directory contains log files containing running information recorded during analysis.
The file name adopts the date it is created. (2016_Dec_06.run.txt)
3-5.Result¶
Result folder
The Result folder stores the output of Octopus-toolkit.
The folder name is based on the GEO accession number you entered. For the private data, the folder name begins with P_.
The Graph folder stores Heatmaps and Lineplots when you run the Graph function.
The detailed information regarding the output can be founded : Output Link
3-7.Tools¶
Tools folder
The Tools directory contains binary versions of 3rd party tools used in the Octopus-toolkit.